
Applied Science Letters


Honeybees play a crucial role in agriculture and the ecosystem through pollination services. They are susceptible to numerous pathogens. Nosemosis is one of the pathological conditions that obvious repercussions on honeybee colonies and is caused by the microsporidium belonging to the genus-Nosemosis. Nosema spp is an obligatory intracellular, ventricular parasite belonging to the kingdom Fungi. The present study focused on the etiology of Nosemosis in honeybee colonies at Malakand Division. Due to the global prevalence of Nosemosis and the huge extermination of honeybee colonies by Nosemosis, we aimed to evaluate Nosemosis in Malakand Division. Random sampling was carried out from the length and breadth of the Division. During sampling, we collect symptomatic older foraging bees and also cadavers near the boxes. We analyzed the sample slides through 400x magnification under a light microscope and used Giemsa stain to improve contrast. We work for about eight months. All of our summer and autumn samples were negative for Nosema spores, but the spring samples were positive for spores. For the prevalence and abundance of spores, we conducted hemocytometry for positive samples and compared the findings with previous literature. The present study highlights that the prevalence rate of Nosemosis is low in the Malakand Division. Further studies are recommended to evaluate Nosemosis through modern molecular techniques, considering all possible sites of the region spanning over the entire year.
INTRODUCTION
Honeybees are one of the most magnificent creatures of nature. They are eukaryotic organisms belonging to the Phylum Arthropoda, having four species, i.e, Apis dorsata (the rock-bee), Apis florea (the little-bee), Apis cerana (Asian honey bee), Apis mellifera (European bee). They play an important role in the economy of the country as they provide honey, Royal jelly, propolis, bee pollen, bee bread, venom, and wax [1], which provides $216 billion to the world economy [2]. Their imparting nature provides many advantages, but along with that, the transmission has negative impacts on the honey bees due to which a number of diseases spread in their population, like brood disease, American foulbrood, European foul-brood, chalk brood, sac brood, Nosema disease, bee paralysis, acarine disease, mites disease, etc [3, 4]. This research is about Nosema disease [5, 6].
Nosema disease or Nosemosis is a disease of adult honey bees, which is caused by a microsporidium-parasite of the genus Nosema and is responsible for significant economic losses for beekeepers-worldwide, acting alone or in combination with other synergistic factors [7]. The genus Nosema has two identified species, namely Nosema apis and Nosema ceranae [8, 9]. They are classified as highly evolved fungi, intracellular obligate parasites of eukaryotes, which are distributed worldwide [10]. N. apis is found in European honeybees (A.mellifera), and N. ceranae was considered to be specific only to the Asian honey bee, but now it is reported that N. ceranae affects the European honeybees as well [10]. It was believed that Apis mellifera are endemically infected by Nosema apis [11]. However, within the last decade, Nosema ceranea that infects the eastern honeybee (Apis ceranae) has been detected in Apis mellifera [12, 13].
Nosema ceranae has been quietly spreading throughout the world’s Aapis mellifera populations since before 1990. According to Klee et al. (2007) and Paxton et al. (2012), it was first discovered in the United States, Brazil, China, Taiwan, Vietnam, Spain, Greece, Italy, Serbia, Germany, France, Denmark, Finland, and Sweden in 2007 [14]. More recently, it was discovered in Canada, Argentina, Hungary, and the United Kingdom. In the UK, Nosema ceranae has been reported in England, Wales, and Northern Ireland, but not in Scotland [15]. In many parts of the world, Nosema ceranae spreads to those areas where Nosema apis already exists [14, 16, 17]. Nosema spores are the main causative agents of Nosema disease [18]. There is a consistent difference between the spores of Nosema ceranae and Nosema apis, but it is difficult to detect the difference in infected bees because mixed infection of both species [19], i.e., Nosema apis and Nosema ceranae, can occur [16] even in individual bees [20]. Both types of spores show variations in shape and size. Spores of Nosema apis have an oval shape, and the spores of Nosema ceranae are often slightly bent, and appear less uniform in shape compared to those of N.apis spores [21]. The size of Nosema ceranae and Nosema apis are 6 x 3 µm and 4.7 x 2.7 µm, respectively. Spores of Nosema ceranae are clearly smaller than those of Nosema apis. Winter is a favorable time for the transmission of Nosema spores due to related stress [22].
The infection spreads through oral-oral or oral-faecal route, contact with aninfected bee, or mating with an infected partner [23]. By ingesting the spores, they begin to develop in the mid gut by protruding a polar filament to deliver the sporoplasm to the epithelium [12]. The epithelial cells burst to release primary spores, which infect the adjacent cells and become secondary or environmental cells to excrete outside. The highest proportion of infected bees of Nosema is found in foraging bees [7]. The infection of Nosema apis is termed Nosemosis type A, and that of Nosema ceranae is Nosemosis type-B. The current research aimed to evaluate Nosemosis and its overall infestation rate in honeybee colonies at Dir Lower Malakand Division (Figure 1), Pakistan.
|
|
|
Figure 1. Map of the study area, Malakand Division, Pakistan |
MATERIALS AND METHODS
Study area
The study was carried out at Malakand and Lower Dir, Pakistan, which are located at 34.00 north and 71.32 ͦ east. The study area has a rich and diverse flora and suitable climatic conditions for honey bee colonies.
Samples collection
Random sampling of infected honey bees (n = 100) was carried out from different managed and wild honey bee colonies in the study area. The honey bee samples were preserved in 70 % ethanol. The abdomens of (n =100) preserved honey bees were dissected and macerated in a mortar and pestle containing 1ml of distilled water per bee. The spore suspension was filtered out through Whatman filter paper remove rough bee parts. Finally, the raw materials trapped were discarded, and the filtrate was then examined under the Light microscope.
Microscopy and micrometry
The isolated spores in the suspensions are placed on clean glass slides for microscopic examination under 400X magnification of a compound microscope for the detection of Nosema spores. Giemsa stain was used improve contrast. Ocular and stage micrometers were used for the calculation of dimensions (length and width) of Nosema spores.
Hemocytometry
A hemocytometer was used for counting the number of spores in different samples. Identification of the Nosema spp (Figure 2). was carried out by using previous literature and taxonomic keys [24].
|
|
|
Figure 2. Nosema spores under 400 x magnification |
RESULTS AND DISCUSSION
In the presence study, samples (n=100) were preserved in 70% ethanol and analyzed under a light microscope (400x magnification). The duration of sampling was 8 months from October 2023 to May 2024. We found that 9 samples out of 100 were positive. The samples that came back positive were collected in the spring season. The ideal sampling time for the identification of Nosema spores is August and September because in these months, the number of spores in the hives increases and reaches the peak in spring, as reported by [25-27].
As the sampling time of the present study is from October to May, which is not an ideal time for Nosema study, the results are not that strong (Table 1). The quality of food available can also be the reason for Nosema development [25], as the autumn flowering plant, such as eucalyptus, yields poor nutrition, which is not ideal for wintering honeybees [25]. As in the Malakand division, there is diverse flora that might be the reason for the low level of Nosemosis. Colony disturbances while moving in winter to new suitable places can also be one of the reasons for Nosema development (Michael Hornitzky), as most of the commercial beekeepers in the Malakand division are migratory and move from one place to another for availing the suitable climatic condition for bees and honey flow, the Nosemosis to Malakand division might have been spread/transfer from another locality [28].
The Nosema spores are transmitted to the neighboring bee through fecal materials and oral-oral exchange. In contrast, the present study didn’t attempt to identify any source of infection; however, the route of spread in the transmission of infective spores may play a role in Nosema transmission [29].
The spores’ count (Table 2) of the positive samples (Table 3) of the present study can be compared with findings of [24] in which the spores count of positive samples were 6.6 x 105, 2.9 x 105, 2.1 x 105, 1.36 x 105, 1 x 105, 3.4 x 105, and 2.7 x 105. The findings (spore count) of the present study show a satisfactory similarity with the work of Bollan et al. (2013) [24].
Table 1. Number of samples and honeybees collected from various areas
|
Collection Area |
Coordinates |
No. of Samples |
No. of Honey bees |
|
Thana |
34.6309° N, 72.0764° E |
25 |
75 |
|
Dargai |
34.1798° N, 71.8883° E |
11 |
20 |
|
Swat |
34.1798° N, 71.8883° E |
10 |
40 |
|
Samarbagh |
34.0506° N, 71.6065° E |
8 |
24 |
|
Kityari |
34.7500° N, 72.0676° E |
17 |
34 |
|
Talash |
34.7415° N, 71.8720° E |
18 |
36 |
|
Timergara |
34.8278° N, 71.8423° E |
11 |
25 |
|
Total |
|
100 |
254 |
Table 2. Nosema spores count for positive samples per honey bee
|
S. No. |
Sample Code |
Number of spores per bee |
|
1 |
1 |
2.6 x106 |
|
2 |
35 |
2.4 x 106 |
|
3 |
36 |
2.9 x 106 |
|
4 |
37 |
2.1 x 106 |
|
5 |
64 |
2.5 x 106 |
Table 3. Apis mellifera collected from various localities and their results under Light microscopy (400x magnifications)
|
Localities |
Spores |
Results |
Sample |
Localities |
Spores |
Results |
|
|
1 |
Lower Dheri |
2.0 |
+ve |
51 |
Kityari |
0 |
-ve |
|
2 |
Upper Dheri |
0 |
-ve |
52 |
Kityari |
0 |
-ve |
|
3 |
Upper Dheri |
0 |
-ve |
53 |
Kityari |
0 |
-ve |
|
4 |
Nari Shah |
0 |
-ve |
54 |
Kityari |
0 |
-ve |
|
5 |
Barikot Kas |
0 |
-ve |
55 |
Kityari |
0 |
-ve |
|
6 |
Barikot Kas |
0 |
-ve |
56 |
Kityari |
0 |
-ve |
|
7 |
Gombat |
0 |
-ve |
57 |
Kityari |
0 |
-ve |
|
8 |
Khalisa |
0 |
-ve |
58 |
Kityari |
0 |
-ve |
|
9 |
Khalisa |
0 |
-ve |
59 |
Kityari |
0 |
-ve |
|
10 |
Bajawro |
0 |
-ve |
60 |
Kityari |
0 |
-ve |
|
11 |
Shoghale |
0 |
-ve |
61 |
Kityari |
0 |
-ve |
|
12 |
Kamartangi |
0 |
-ve |
62 |
Kityari |
0 |
-ve |
|
13 |
ChamBajawro |
0 |
-ve |
63 |
Kityari |
0 |
-ve |
|
14 |
Chino |
0 |
-ve |
64 |
Kityari |
1.9 |
-ve |
|
15 |
Nasafa |
0 |
-ve |
65 |
Kityari |
0 |
-ve |
|
16 |
Baldara |
0 |
-ve |
66 |
Badam baghicha |
0 |
-ve |
|
17 |
Tekas |
0 |
-ve |
67 |
Gul baba |
0 |
-ve |
|
18 |
Nari Shah |
0 |
-ve |
68 |
Ghora chock |
0 |
-ve |
|
19 |
Samarbagh |
0 |
-ve |
69 |
Ghora chock |
0 |
-ve |
|
20 |
Samarbagh |
0 |
-ve |
70 |
Batram |
0 |
-ve |
|
21 |
Samarbagh |
0 |
-ve |
71 |
Maizara |
0 |
-ve |
|
22 |
Samarbagh |
0 |
-ve |
72 |
Maizara |
0 |
-ve |
|
23 |
Samarbagh |
0 |
-ve |
73 |
Maizara |
0 |
-ve |
|
24 |
Samarbagh |
0 |
-ve |
74 |
Ghani Khan Road |
2.4 |
+ve |
|
25 |
Samarbagh |
0 |
-ve |
75 |
Ghani Khan Road |
0 |
-ve |
|
26 |
Samarbagh |
0 |
-ve |
76 |
Ghani Khan Road |
2 |
-ve |
|
27 |
Timegara |
0 |
-ve |
77 |
Chakdarra |
0 |
-ve |
|
28 |
Timegara |
0 |
-ve |
78 |
Dandha |
0 |
-ve |
|
29 |
Timegara |
0 |
-ve |
79 |
Dandha |
0 |
-ve |
|
30 |
Timegara |
0 |
-ve |
80 |
Dandha |
0 |
-ve |
|
31 |
Timegara |
0 |
-ve |
81 |
Complex |
0 |
-ve |
|
32 |
Timegara |
0 |
-ve |
82 |
Complex |
0 |
-ve |
|
33 |
Timegara |
0 |
-ve |
83 |
Thana |
0 |
-ve |
|
34 |
Timegara |
0 |
-ve |
84 |
Complex |
0 |
-ve |
|
35 |
TimegaraTrai |
1.8 |
+ve |
85 |
Lal colony |
0 |
-ve |
|
36 |
TimegaraTrai |
2.2 |
+ve |
86 |
Lal colony |
0 |
-ve |
|
37 |
TimegaraTrai |
1.6 |
+ve |
87 |
Amjadshaheed Colony |
0 |
-ve |
|
38 |
KoatManzary |
0 |
-ve |
88 |
Sunehri Masjid |
0 |
-ve |
|
39 |
KoatManzary |
0 |
-ve |
89 |
Khadimabad |
0 |
-ve |
|
40 |
KoatManzary |
0 |
-ve |
90 |
Khadimabad |
0 |
-ve |
|
41 |
Jabanr |
0 |
-ve |
91 |
Swat |
0 |
-ve |
|
42 |
Jabanr |
0 |
-ve |
92 |
Swat |
0 |
-ve |
|
43 |
Jabanr |
0 |
-ve |
93 |
Swat |
0 |
-ve |
|
44 |
Dargai |
0 |
-ve |
94 |
Swat |
2.1 |
+ve |
|
45 |
Dargai |
0 |
-ve |
95 |
Swat |
0 |
-ve |
|
46 |
Dargai |
0 |
-ve |
96 |
Swat |
0 |
-ve |
|
47 |
Dargai |
0 |
-ve |
97 |
Swat |
2.3 |
+ve |
|
48 |
Dargai |
0 |
-ve |
98 |
Swat |
2.1 |
+ve |
|
49 |
Kityari |
0 |
-ve |
99 |
Swat |
0 |
-ve |
|
50 |
Kityari |
0 |
-ve |
100 |
Swat |
0 |
-ve |
Light microscopy revealed that fresh Nosema ceranae spores are oval and round in shape, varying in 124 size (Figure 3). The images are given below, indicating the spores of Nosema ceranae.
|
|
|
Figure 3. Counting of Nosema spores via hemocytometer |
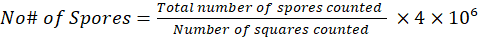
|
|
(1) |
CONCLUSION
In conclusion, this analysis underscores the heterogeneous distribution of fungal spores across different localities, highlighting both areas of potential concern and those less impacted. These findings provide crucial insights for local health authorities and policymakers to prioritize monitoring and mitigation efforts aimed at reducing potential health risks associated with fungal spores in specific regions. The accurate identification of Nosema spores to spp level by molecular techniques should be recommended. Research should be conducted for the treatment of Nosemosis. Integrated pest management strategies should be improved for honey bee health. Research grants in apiculture should be encouraged by the government.
ACKNOWLEDGMENTS: Department of Zoology, University of Malakand, is highly acknowledged for providing support.
CONFLICT OF INTEREST: None
FINANCIAL SUPPORT: None
ETHICS STATEMENT: The animal study was reviewed and approved by the ethical committee of the Department of Zoology, University of Malakand with (Ref. no. E-SA-11-2009).
